For decades, lipid biology has had a frustrating blind spot: we knew lipids were doing important things inside cells — binding to proteins, organizing membranes, regulating signaling — but we had no way to catch them in the act inside a living cell. Most techniques required breaking the cell open first, which destroys the very context you're trying to study.
A new paper in Nature Cell Biology (May 2026) just changed that.
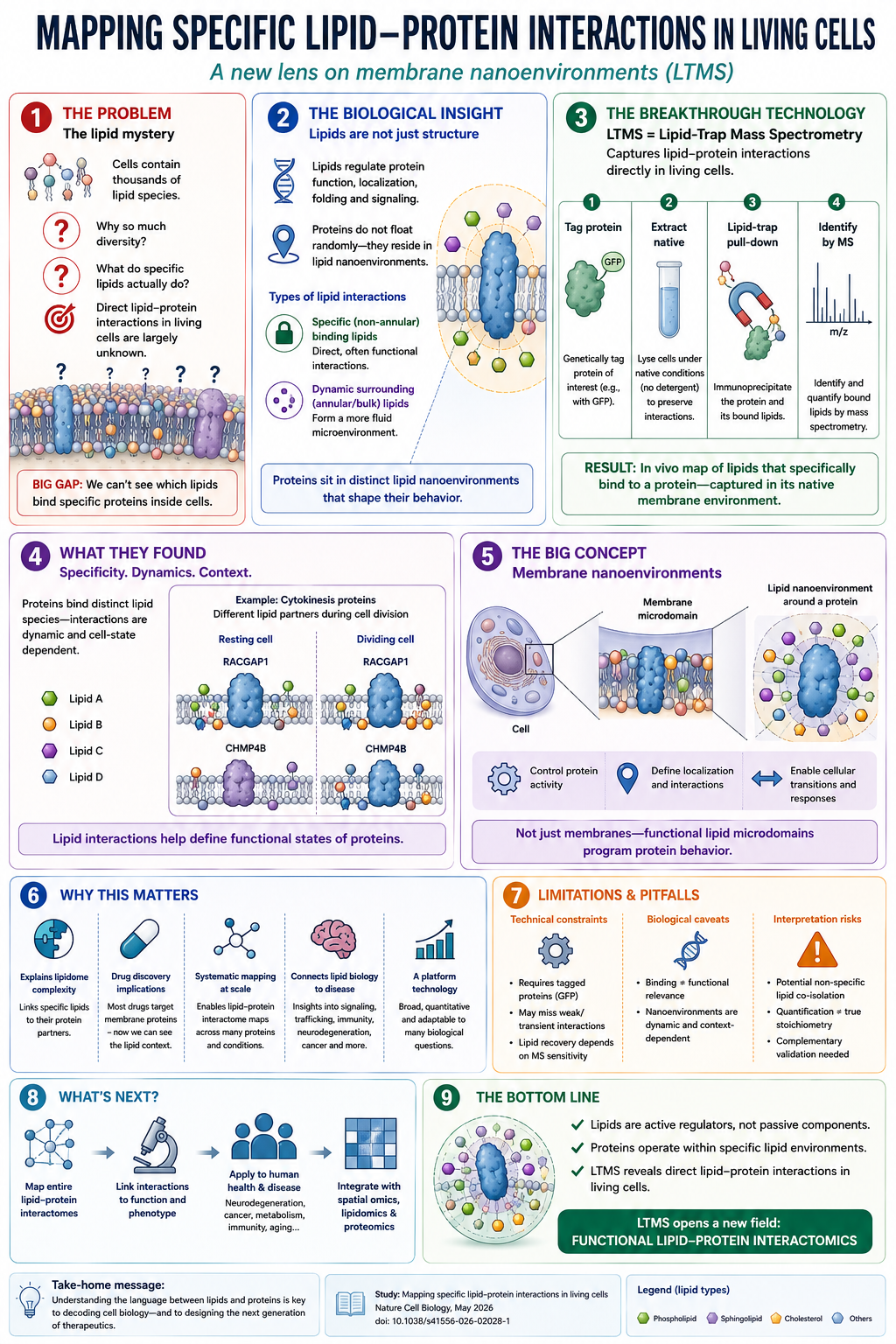
The New Tool: Lipid-Trap Mass Spectrometry
Researchers at King's College London developed a technique called lipid-trap mass spectrometry (LTMS) — a method to systematically identify which lipids are directly bound to specific proteins inside living mammalian cells.
The approach is elegant: tag a protein of interest with a fluorescent marker (GFP), then pull it out of the cell using immunoprecipitation — a well-established technique that "fishes out" tagged proteins. But instead of just analyzing the protein, also analyze the lipids that came along for the ride. Mass spectrometry then identifies precisely which lipids were bound to the protein at the moment of capture.
Paquola A, Benson CE, Desale SE et al. "Lipid-trap mass spectrometry identifies lipid–protein interactions in cells." Nature Cell Biology, 2026. DOI: 10.1038/s41556-026-01928-6
What They Found
The team applied LTMS to cell division — one of the most precisely regulated biological processes — and found something striking: specific proteins involved in cell division bind to specific lipid species in dividing cells that they don't bind in non-dividing cells. The lipid environment around a protein is not random. Proteins appear to operate in selective lipid nanoenvironments — tiny membrane patches with a specific lipid composition that may be essential for the protein to function correctly.
This has profound implications. If a protein needs a particular lipid to function, then disrupting that lipid — through disease, diet, or a drug — could directly impair the protein's activity without touching the protein itself.
Why This Matters for Disease
Almost every major disease — Alzheimer's, cancer, metabolic syndrome, neurodegeneration — is associated with changes in cellular lipid composition. We've known this for years. What we haven't been able to do is connect specific lipid changes to specific protein dysfunctions in living cells.
LTMS is the tool that makes that connection possible. It opens a completely new approach to understanding how lipid changes drive disease at the molecular level — and potentially, how targeting specific lipid-protein interactions could become a new class of therapeutics.
As Professor Eggert, the paper's senior author, put it: the goal is understanding what lipids actually do in cells — not just where they are. LTMS is a major step toward that answer.
Limitations and Pitfalls
LTMS is a powerful new tool, but like all new technologies it has real constraints worth understanding:
What's Next
Lipids are active regulators of cell biology, not passive structural components. Proteins operate within specific lipid environments that shape their function. LTMS now lets us map those environments directly in living cells. This opens a new field: Functional Lipid-Protein Interactomics. The implications for drug discovery — designing molecules that target not just proteins, but the lipid context those proteins require — are only beginning to be understood.